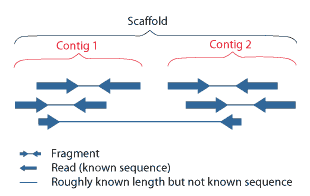
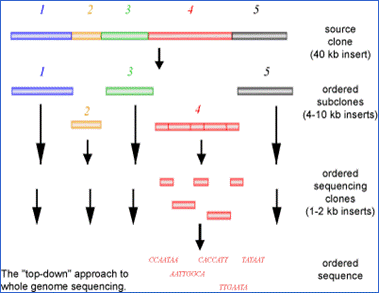
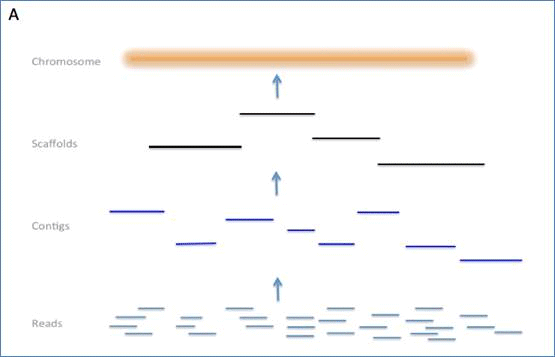
Clone Contig Sequencing Methods | Clone Contig Approach | Whole Genome Sequencing By Clone Contig | - YouTube
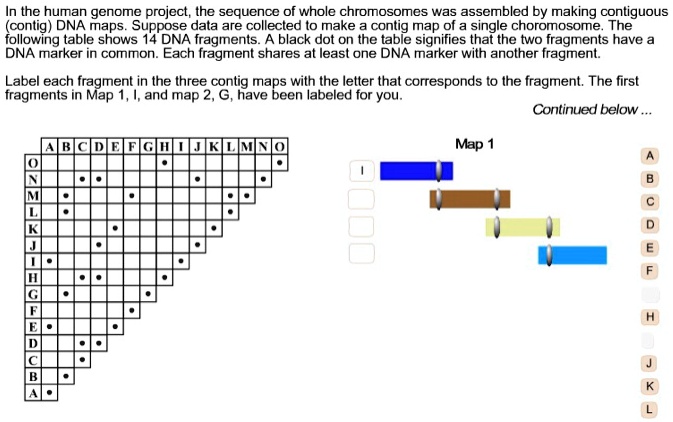
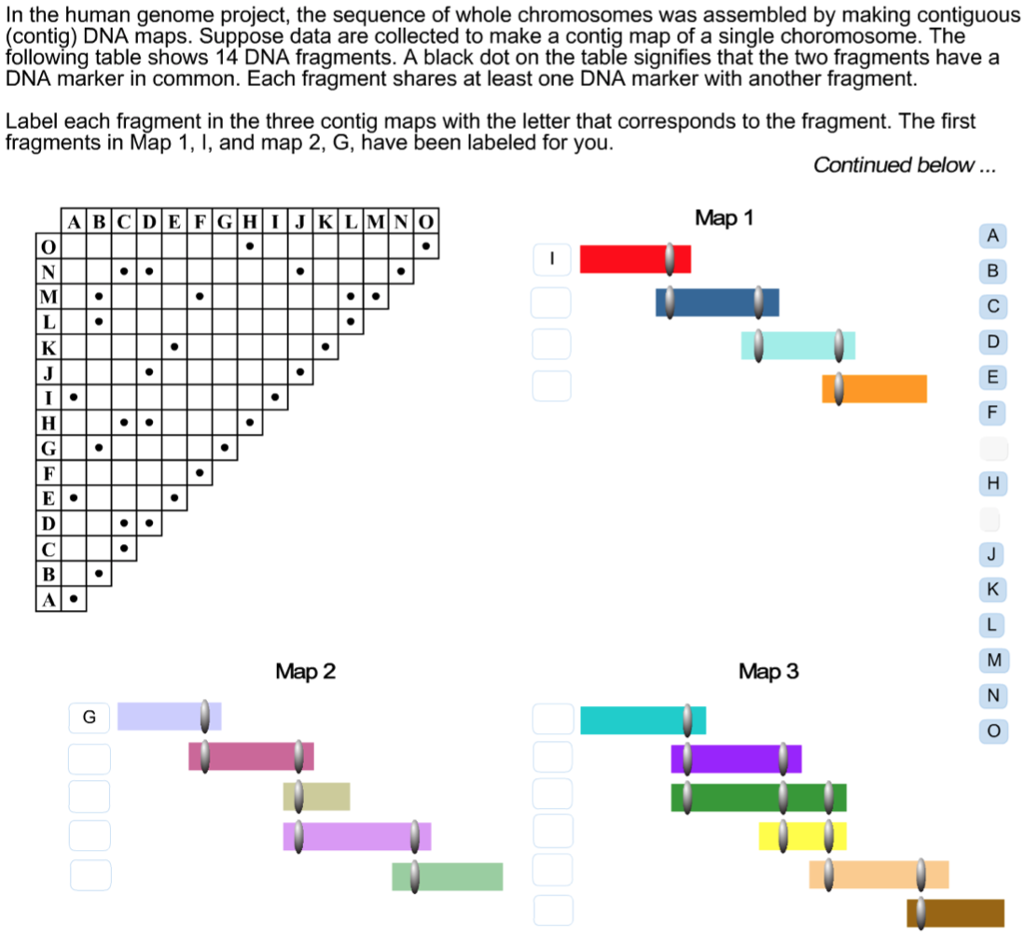
SOLVED: In the human genome project; the sequence of whole chromosomes was assembled by making contiguous contig DNA maps. data are collected make contig map of single choromosome The Suppose following table
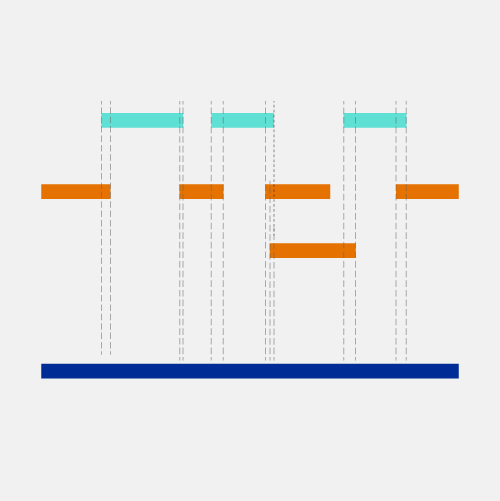
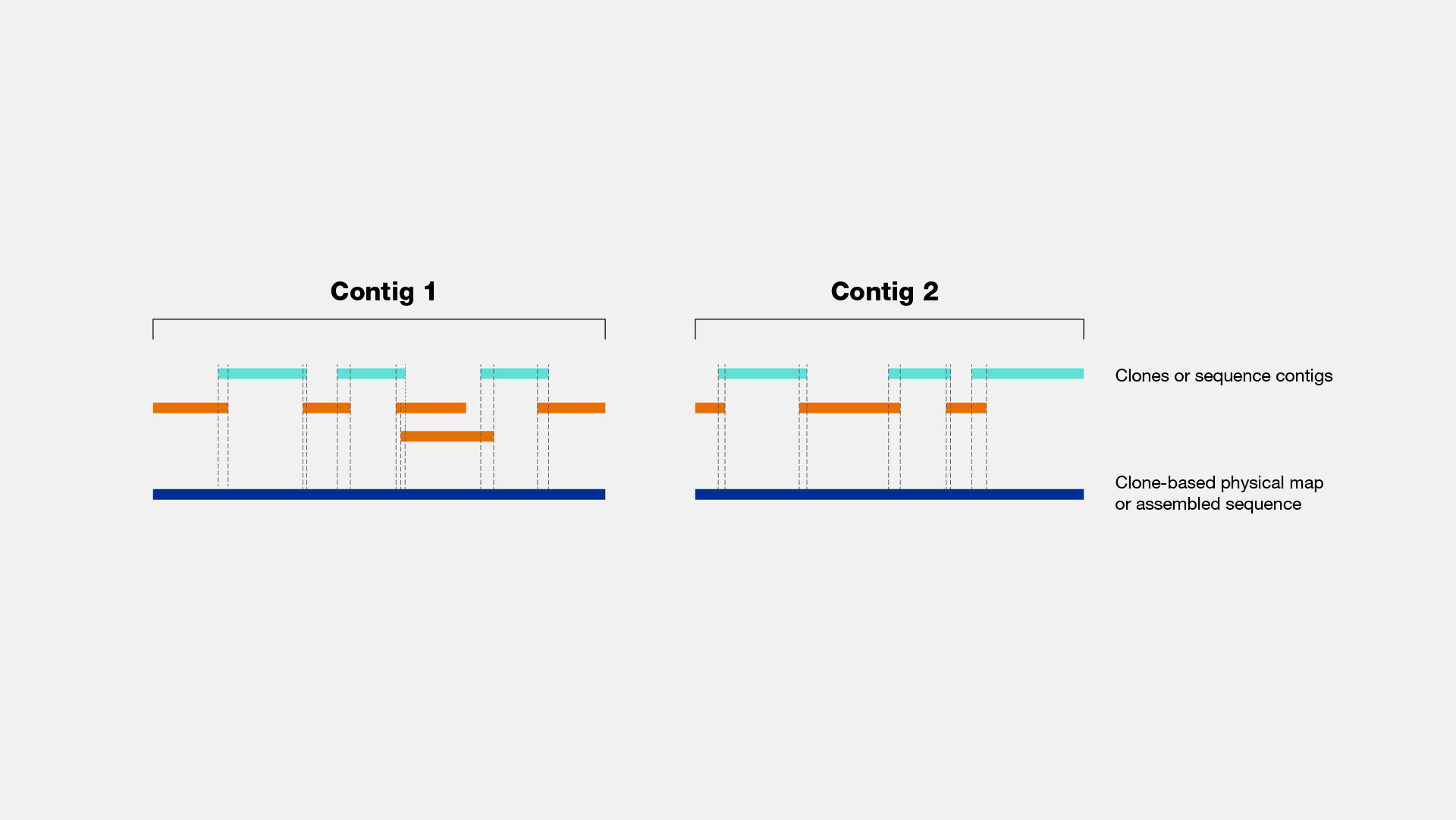
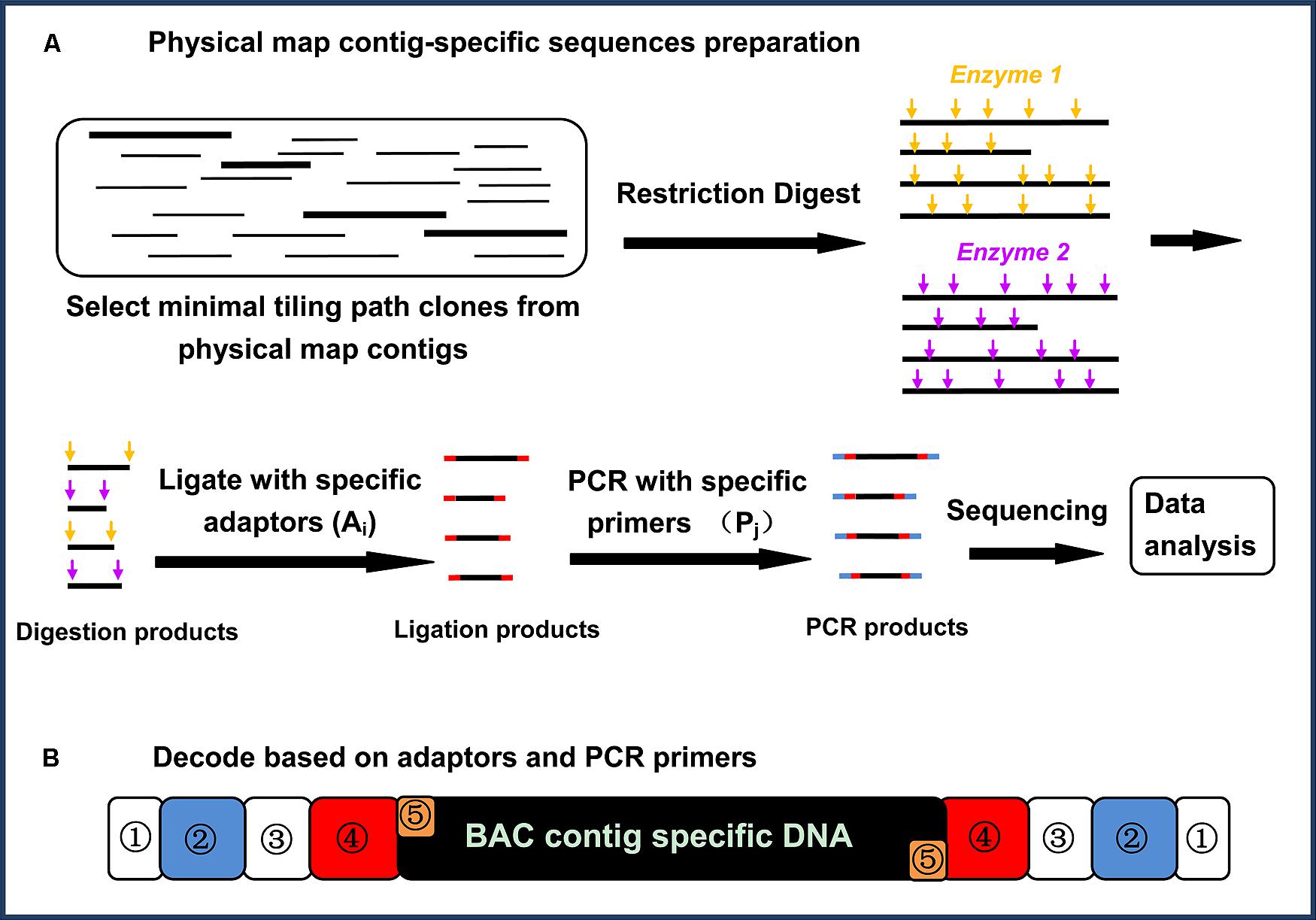
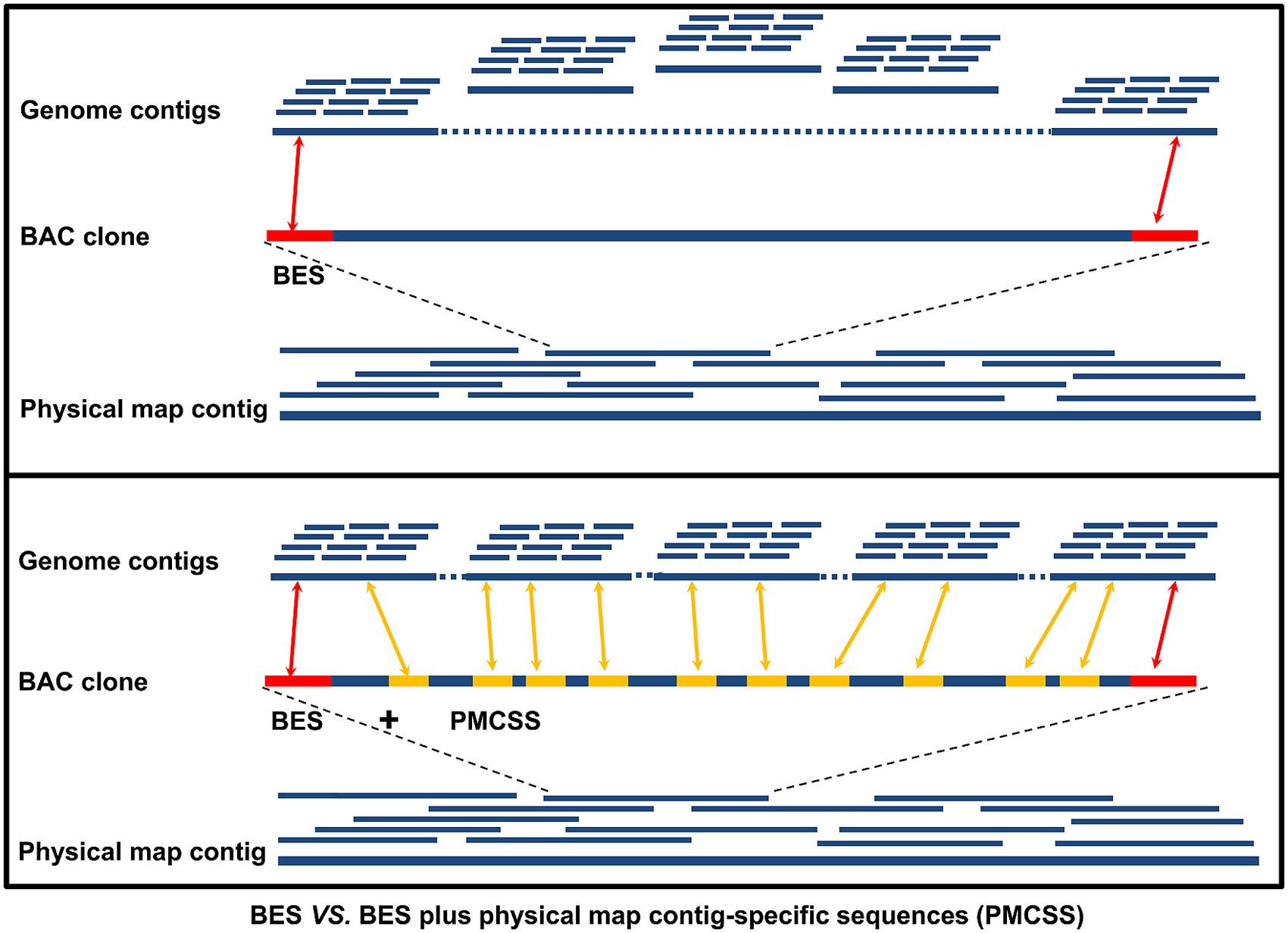
Generation of Physical Map Contig-Specific Sequences Useful for Whole Genome Sequence Scaffolding | PLOS ONE

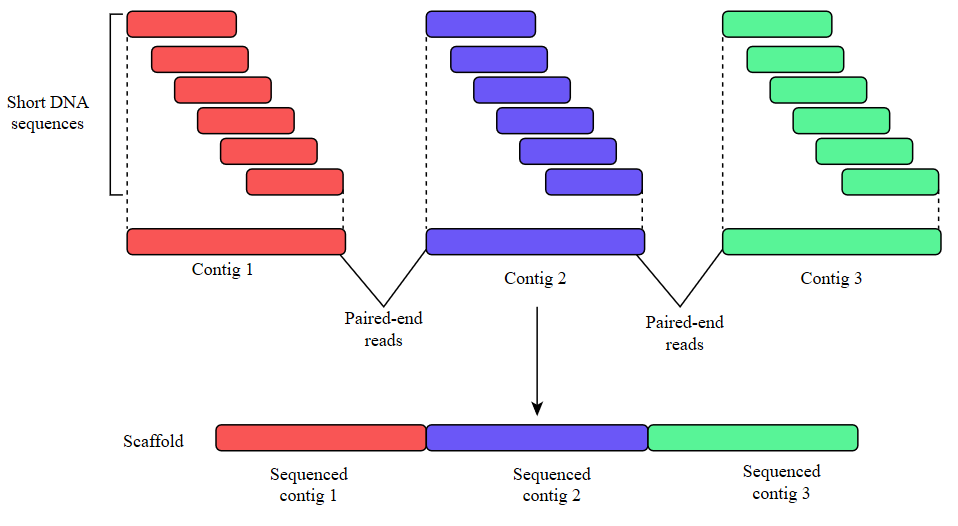
Whole Genome Sequencing Process of Creating Contigs from Fragment Reads | Download Scientific Diagram

Introduction to the principles and methods underlying the recovery of metagenome‐assembled genomes from metagenomic data - Goussarov - 2022 - MicrobiologyOpen - Wiley Online Library
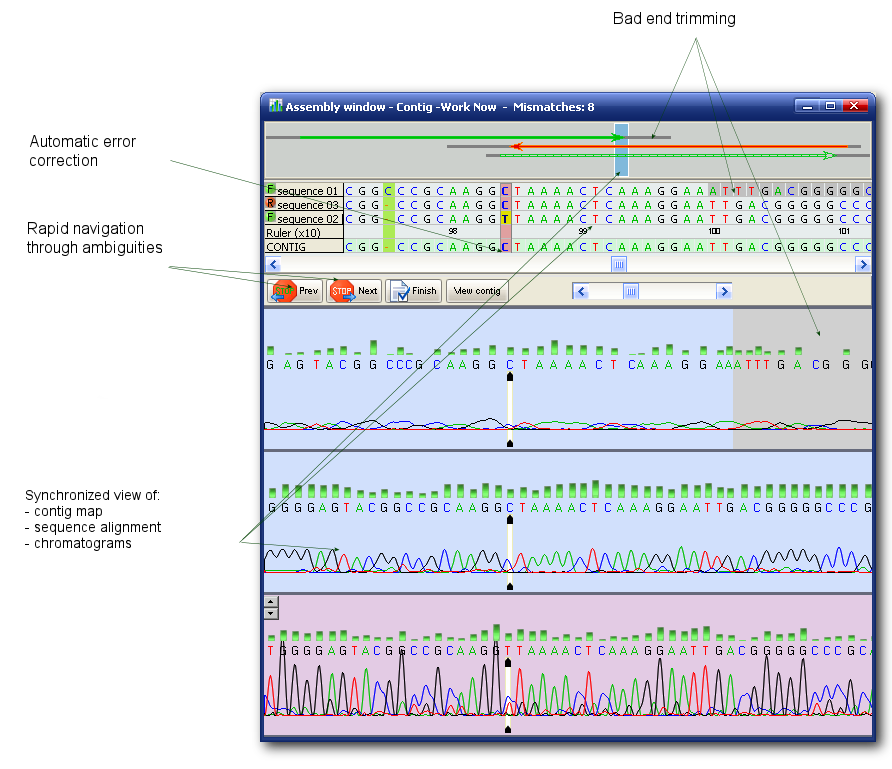
DNA Sequence Assembler->Manual->Contig Assembly window|The contig map, contig grid, reference and chromatogram panel. An affordable alternative to the really expensive software for DNA assembly and alignment on the market. DNA BASER it
![PyDamage: automated ancient damage identification and estimation for contigs in ancient DNA de novo assembly [PeerJ] PyDamage: automated ancient damage identification and estimation for contigs in ancient DNA de novo assembly [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/11845/1/fig-1-full.png)